Random Numbers#
Start with NumPy and matplotlib:
import numpy as np
import matplotlib.pylab as plt
Random Number Generation#
Get a random number via NumPy’s random module, using its rand() function:
a = np.random.rand()
print( a )
0.7672451703800369
Get a whole random matrix:
A = np.random.rand(3, 3, 2)
print( A )
[[[0.99227986 0.30622325]
[0.30131488 0.59967493]
[0.63837998 0.54534001]]
[[0.62730555 0.94456943]
[0.55941718 0.71779347]
[0.46527538 0.45750565]]
[[0.37142985 0.87645323]
[0.3180609 0.80357048]
[0.24840259 0.15517278]]]
True vs. Pseudo Random Numbers#

Computer algorithms commonly create pseudo random numbers with “good” statistics, but being dependent on a seed:
... 93802948542771471902796694497278419429380485689050661139408044323 ...
| start here ... | or here ...
in NumPy we can set the seed on our own:
np.random.seed(29)
a = np.random.rand()
print( a )
0.8637599855700829
which will set the whole series of following random numbers:
np.random.seed(29)
a = np.random.rand()
print( a )
a = np.random.rand()
print( a )
a = np.random.rand()
print( a )
0.8637599855700829
0.2849059652276268
0.07325638808373602
Note
Seeding can be handy to test algorithms using random numbers!
Different Statistics#
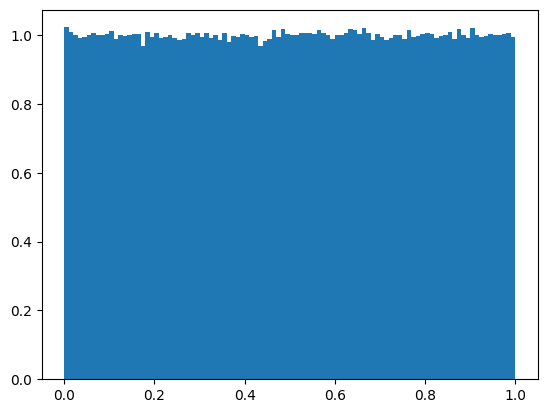
np.random.rand(): Uniform distribution#
Get a random vector and plot its histogram:
N = 1000000
b = np.random.rand(N)
plt.figure(1)
plt.hist(b, bins=100, density=True)
plt.show()
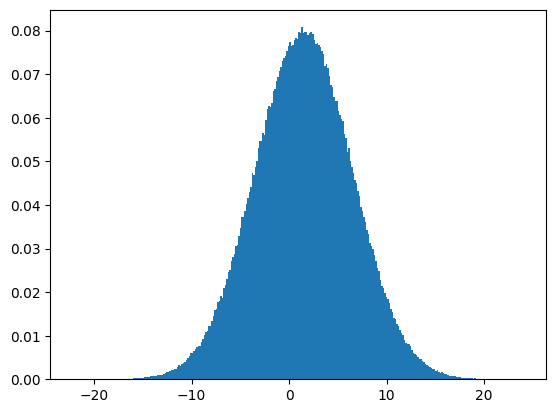
np.random.normal(): Normal (Gaussian) distribution#
N = 1000000
b = np.random.normal(size=N, loc=1.5, scale=5.0)
plt.figure(1)
plt.hist(b, bins=300, density=True)
plt.show()
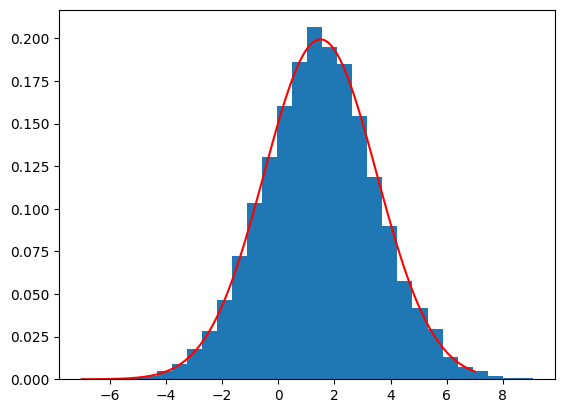
#?np.random.normal
def gauss(mu, sigma, x):
return 1/(sigma * np.sqrt(2 * np.pi)) \
* np.exp( - (x - mu)**2 / (2 * sigma**2) )
plt.figure(1)
b = np.random.normal(size=10000, loc=1.5, scale=2.0)
plt.hist(b, bins=30, density=True)
xPlot = np.linspace(-7, 7, 100)
plt.plot(xPlot, gauss(mu=1.5, sigma=2.0, x=xPlot), 'r-')
plt.show()
Integration via Random Numbers / Coordinates#
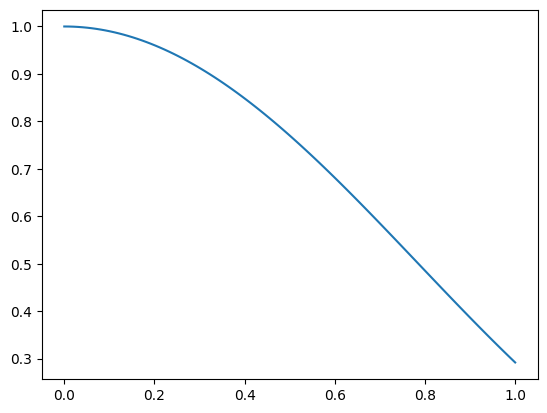
def f(x):
return np.cos(x)**2.0
xFine = np.linspace(0, 1, 100)
plt.figure(1)
plt.plot(xFine, f(xFine))
plt.show()
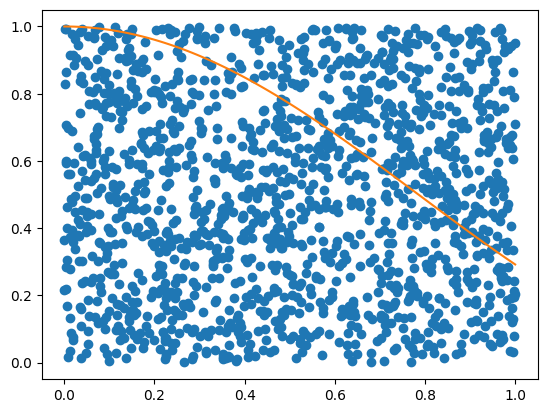
Get random x and y coordinates:
low = [0,0]
high = [1,1]
points = np.random.uniform(low ,high, size=(1500,2))
x, y = points[:,0], points[:,1]
plt.figure(1)
plt.plot(x, y, 'o')
plt.plot(xFine, f(xFine))
plt.show()
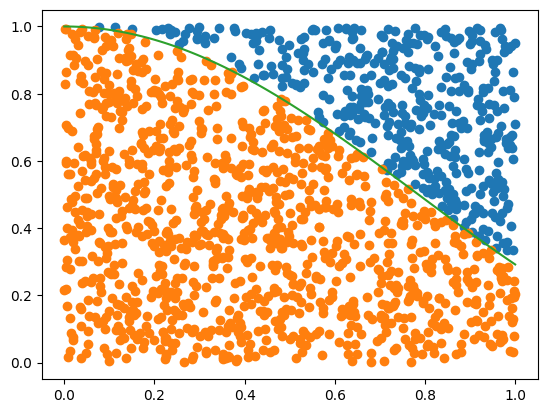
Now mark points below and above f(x):
indAbove = np.where( y >= f(x) )[0]
indBelow = np.where( y < f(x) )[0]
plt.figure(1)
plt.plot(x[indAbove], y[indAbove], 'o')
plt.plot(x[indBelow], y[indBelow], 'o')
plt.plot(xFine, f(xFine))
plt.show()
Finally get an estimate of the integral of f(x) from 0 to 1 by comparing the amount of points below f(x) to the total number of points and compare to SciPy’s quad():
intRand = len(indBelow) / ( len(indBelow) + len(indAbove) )
print("Monte Carlo:", intRand )
import scipy.integrate as integrate
intQuad, error = integrate.quad(f, a=0, b=1)
print(" SciPy:", intQuad )
Monte Carlo: 0.7193333333333334
SciPy: 0.7273243567064205
Multi-Dimensional Integration via Random Numbers / Coordinates#
def f(x,t):
return np.exp(-x*t) / (t**2 + 1)
# get random numbers
N = 1500
x, t = np.random.rand(2, N) * 2.0
y = np.random.rand(N, N) * 2.0
# get the volume of the FULL space we have been sampling
volume = 2.0 * 2.0 * 2.0
# create a meshgrid of our random (x,t) coordinates
# so that we can call f(xGrid, tGrid)
xGrid, tGrid = np.meshgrid(x, t)
# count how often our random y(x,t) lays below f(x,t)
indBelow = np.where( y <= f(xGrid, tGrid) )
# and calculate the interal with the proper normalization!
intRand = len(indBelow[0]) * volume / N**2.0
# multi-dimensional integral via scipy's integrate.nquad
intQuad, error = integrate.nquad(f, [[0, 2],
[0, 2]])
print("Monte Carlo:", intRand )
print(" SciPy:", intQuad )
Monte Carlo: 1.2833422222222222
SciPy: 1.3039147247360718
Mean Values via Random Numbers#
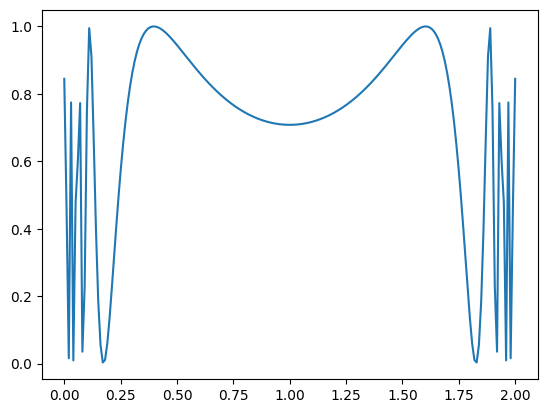
for $\( f(x) = sin \left( \frac{ 1 }{x(2-x)} \right)^2, \)\( which is problematic for \)a=0\( and \)b=2$.
# define a function f(x) we like to consider
def f(x):
return (np.sin(1/(x*(2-x))))**2
N = 200
x = np.linspace(0.0001, 1.9999, N)
y = f(x)
plt.figure(1)
plt.plot(x, y)
plt.show()
#?integrate.quad
d = 0.1
a = 0 + d
b = 2 -d
intQuad, error = integrate.quad(f, a=a, b=b)
print( 1/(b-a) * intQuad )
0.7550327874373094
Let’s calculate the standard mean for a variety of starting points x0:
N = 10000
for x0 in np.logspace(-1, -6, 8):
print( "%f : %f"%(x0, f(np.linspace(x0, 1, N)).mean() ))
0.100000 : 0.755029
0.019307 : 0.729971
0.003728 : 0.726605
0.000720 : 0.725740
0.000139 : 0.726149
0.000027 : 0.725691
0.000005 : 0.725232
0.000001 : 0.725425
And for a variety of N:
for N in range(100000, 1000000, 100000):
print( "%d : %f"%(N, f(2.0 * np.random.rand(N)).mean()))
100000 : 0.725837
200000 : 0.726622
300000 : 0.725721
400000 : 0.726297
500000 : 0.725663
600000 : 0.725421
700000 : 0.725388
800000 : 0.725737
900000 : 0.726596
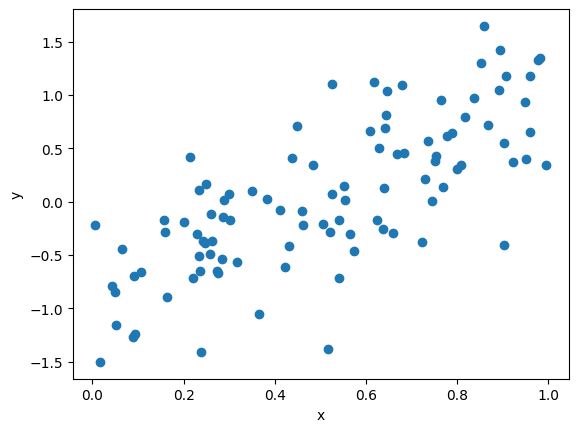
Simulating an Experiment via Random Numbers#
N = 100
# create x on a random grid
x = np.random.rand(N)
# and y = 2.0 * x - 1.0
# ... and add Gaussian noise (like in an experiment) to it
y = 2.0 * x - 1.0 + np.random.normal(scale=0.5, size=N)
# and plot everything
plt.figure(1)
plt.plot(x, y, 'o')
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# load curve_fit from scipy
from scipy.optimize import curve_fit
# define a linear function
def fitFunc(x, a, b):
return a * x + b
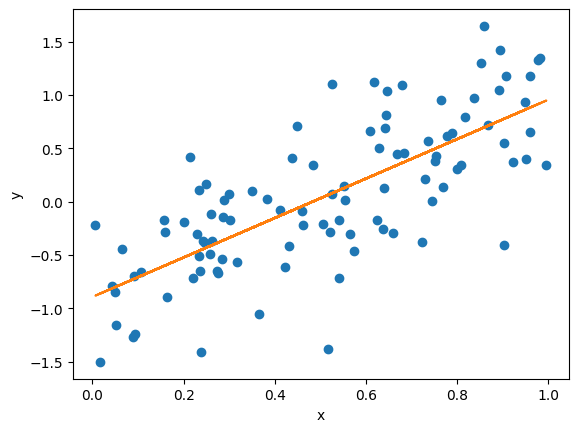
# and fit it to the noisy data
myFit, V = curve_fit(fitFunc, x, y)
# get error estimate
error = np.sqrt(np.diag(V))
# print out fitting parameters
print(" a = %+4.3f +/- %4.3f"%(myFit[0], error[0]))
print(" b = %+4.3f +/- %4.3f"%(myFit[1], error[1]))
# and plot everything
plt.figure(1)
plt.plot(x, y, 'o')
plt.plot(x, fitFunc(x, *myFit))
plt.xlabel('x')
plt.ylabel('y')
plt.show()
a = +1.848 +/- 0.168
b = -0.893 +/- 0.099
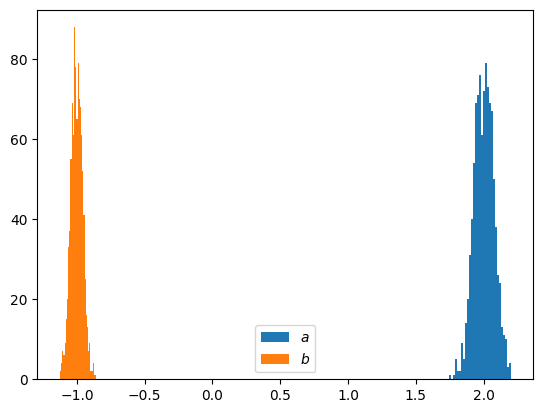
# let's have a look into the STATISTICS of our fitting parameters
N = 1000 # Number of random sets ... for the statistics
a = np.zeros(N) # array to save all fitted a
b = np.zeros(N) # array to save all fitted a
# loop over sets
for i in range(N):
# create a new set
x = np.random.rand(50)
y = 2.0 * x - 1.0 + np.random.normal(scale=0.15, size=50)
# and fit it to the noisy data
myFit, V = curve_fit(fitFunc, x, y)
# get error estimate
error = np.sqrt(np.diag(V))
# save a and b
a[i] = myFit[0]
b[i] = myFit[1]
#plt.subplot(2, 1, 1)
plt.hist(a, bins=int(np.sqrt(N)), label='$a$' )
plt.hist(b, bins=int(np.sqrt(N)), label='$b$' )
plt.legend()
#plt.subplot(2,1,2)
#plt.hist(b, bins=int(np.sqrt(N)), label='$b$' )
#plt.legend()
plt.show()
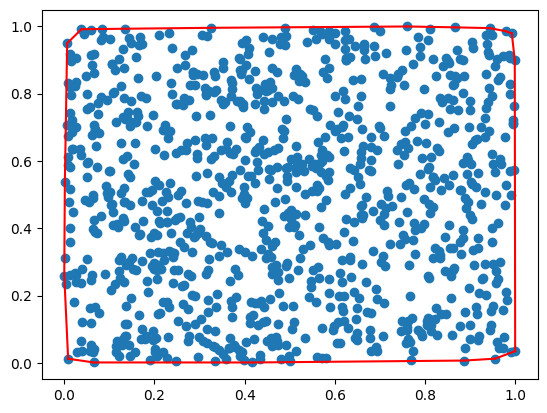
Convex hull of a square#
# function that returns all points of A (set of x/y points)
# which belong to its convex hull
def getHull(A):
N = np.size(A[:,0])
# create an empty list of corner indices
ind = []
# add the top-most (largest y position) point to the list of corner indices
# ... since it belongs to our convex hull by definition
ind.append( np.argmax(A[:,1]) )
# initial values for our search
lastAngle = -4.0
closed = False
# search for all points of the convex hull via while loop
# ... continue the search as long as the hull is not closed
while (not closed):
# Algorithm idea: find the next point with smallest angle
# larger than previous angle
# generate a list of angles between our last point and all other points
newAngles = np.arctan2(A[:, 1] - A[ind[-1], 1],
A[:, 0] - A[ind[-1], 0])
# "penalty" for angle = 0
newAngles[np.where(newAngles == 0)] += 1000.0
# find the smallest angle from our "penalized" angle list
nextInd = np.argmin(np.abs(lastAngle - newAngles))
# check if the next index is already in our list
if(nextInd in ind):
# ... if yes the hull is closed
closed = True
else:
# ... if not we save the angle between the last and the next
# angle as our "lastAngle"
lastAngle = np.arctan2(A[nextInd,1] - A[ind[-1],1],
A[nextInd,0] - A[ind[-1],0])
# and add the nextIndex to the list of our hull array
ind.append( nextInd )
return ind
# generate N random (x,y)
N = 1000
A = np.random.rand(N, 2)
# and plot them
plt.plot(A[:,0], A[:,1], 'o')
ind = getHull(A)
# plot line connecting all corner indices
for ii in range(np.size(ind)):
plt.plot( A[[ind[ii-1], ind[ii]], 0],
A[[ind[ii-1], ind[ii]], 1], 'r')
plt.show()
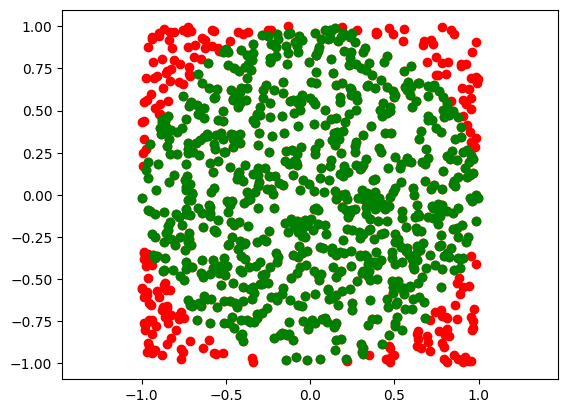
Convex hull of a circle#
# generate N random (x,y)
N = 1000
A = np.random.rand(N, 2) * 2.0 - 1.0
# get all points which belong to a circle of with radius = 1
ind = np.where( np.sqrt( A[:,0]**2.0 + A[:,1]**2.0) <= 1 )[0]
# and plot them
plt.plot(A[:,0], A[:,1], 'or')
plt.plot(A[ind,0], A[ind,1], 'og')
plt.axis('equal')
plt.show()
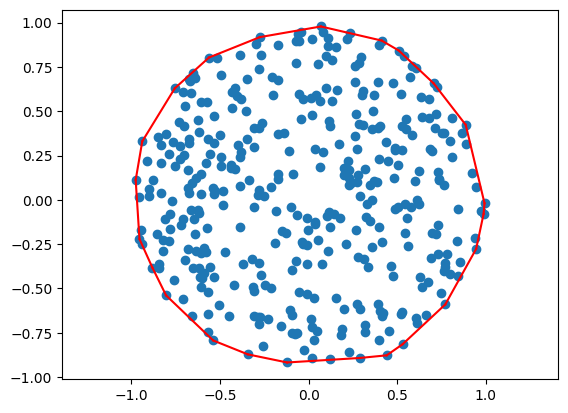
# generate N random (x,y)
N = 500
A = np.random.rand(N, 2) * 2.0 - 1.0
ind = np.where( np.sqrt( A[:,0]**2.0 + A[:,1]**2.0) <= 1 )[0]
# and plot them
plt.plot(A[ind,0], A[ind,1], 'o')
circle = A[ind,:]
ind = getHull(circle)
# plot line connecting all corner indices
for ii in range(np.size(ind)):
plt.plot( circle[[ind[ii-1], ind[ii]], 0],
circle[[ind[ii-1], ind[ii]], 1], 'r')
plt.axis('equal')
plt.show()