Fitting via Optimization#
import numpy as np
import matplotlib.pyplot as plt
import scipy.optimize as optimize
Fitting Simple Functions#
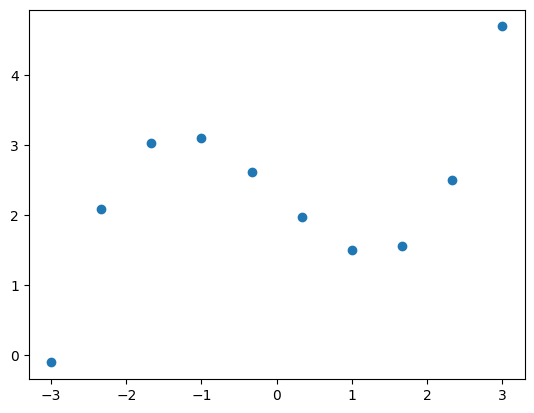
xi = np.linspace(-3, 3, 10)
yi = 0.2 * xi**3.0 - 1.0 * xi + 2.3
plt.figure(1)
plt.plot(xi, yi, 'o')
plt.show()
In the following we define fit(x,c) as our generic fitting function. To find the optimal fitting parameters c (a list here), we further define a cost function cost(c), which measures the quality of the fit by calculating
$\(
cost(\vec{c}) = \sum_i |y_i - f(x_i, \vec{c} )|^2,
\)\(
where \)y_i\( is the original data and \)f(x_i, \vec{c})$ our fit.
Afterwards we use optimize.minimize() to minimize the cost function via finiding the optimal parameters c. With the chosen cost function, this equivalent to a least-square fit.
def fit(x, c):
return c[0] * x**3.0 + c[1] * x + c[2]
#return c[0] * np.cos(x - c[1]) + c[2] * x + c[3]
def cost(c):
return np.sum( np.abs( yi - fit(xi, c) )**2.0 )
sol = optimize.minimize(fun=cost, x0=[0, 0, 0, 0],
method='SLSQP')
print(sol)
message: Optimization terminated successfully
success: True
status: 0
fun: 5.181077947482736e-13
x: [ 2.000e-01 -1.000e+00 2.300e+00 0.000e+00]
nit: 5
jac: [ 5.241e-09 8.601e-10 1.932e-11 0.000e+00]
nfev: 32
njev: 5
multipliers: []
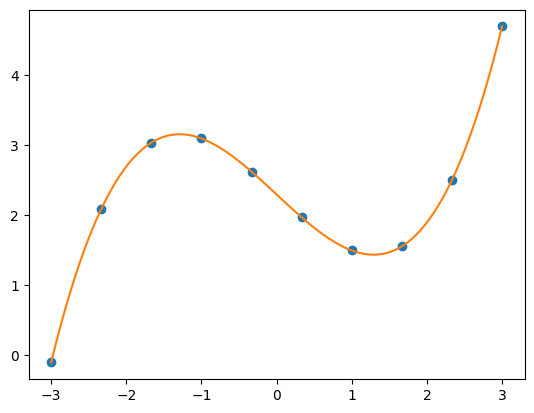
xPlot = np.linspace(-3, 3, 100)
plt.figure(1)
plt.plot(xi, yi, 'o')
plt.plot(xPlot, fit(xPlot, sol.x))
plt.show()
Fitting Noisy Data#
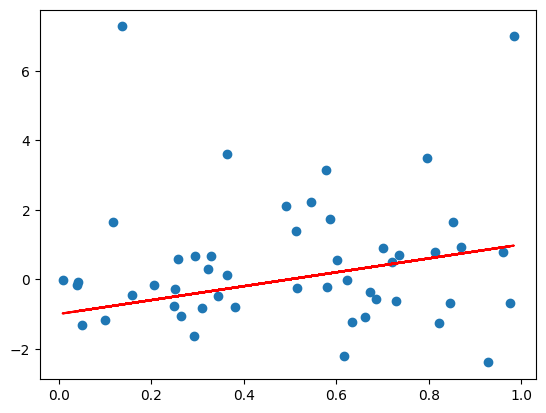
n = 50
x = np.random.rand(n)
# heavly tailed random errors
y_clean = 2.0*x - 1.0
y = 2.0*x - 1.0 + np.random.standard_t(df=2, size=n)
plt.figure(1)
plt.plot(x, y, 'o')
plt.plot(x, y_clean, 'r-')
plt.show()
def fit(x, c):
return c[0] * x + c[1]
# least-square cost function
def costL2(c, xi, yi):
return np.sum( np.abs( yi - fit(xi, c) )**2.0 )
# absolute error cost function
def costL1(c, xi, yi):
return np.sum( np.abs( (yi - fit(xi, c)) ) )
n = 50
N = 1000
errL2 = np.zeros(N)
errL1 = np.zeros(N)
for i in range(N):
x = np.random.rand(n)
y = 2.0*x - 1.0 + np.random.standard_t(df=2, size=n)
solL2 = optimize.minimize(fun=costL2, x0=[0, 0],
method='SLSQP', args=(x, y))
errL2[i] = np.abs(solL2.x[0] - 2)
solL1 = optimize.minimize(fun=costL1, x0=[0, 0],
method='SLSQP', args=(x, y))
errL1[i] = np.abs(solL1.x[0] - 2)
print(" L2 mean:", np.mean(errL2))
print(" L1 mean:", np.mean(errL1))
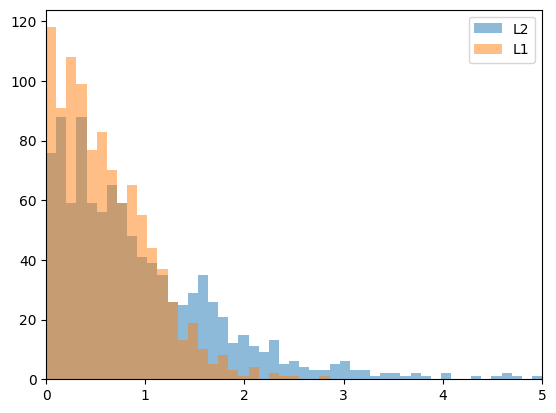
plt.figure(1)
plt.hist(errL2, bins=np.linspace(0,5,50),
label='L2', alpha=0.5)
plt.hist(errL1, bins=np.linspace(0,5,50),
label='L1', alpha=0.5)
plt.xlim([0, 5])
plt.legend()
plt.show()
L2 mean: 1.0278534128251526
L1 mean: 0.6015541731084001
def fit(x, c):
return c[0] * x + c[1]
# least-square cost function
def costL2(c, xi, yi):
return np.sum( np.abs( yi - fit(xi, c) )**2.0 )
# absolute error cost funciton
def costL1(c, xi, yi):
return np.sum( np.sqrt( (yi - fit(xi, c))**2.0 ) )
# logarithmic error cost function
def costLLog(c, xi, yi):
return np.sum(np.log(( fit(xi, c) - yi)**2.0 + 1.0))
n = 50
N = 1000
errL2 = np.zeros(N)
errL1 = np.zeros(N)
errLLog = np.zeros(N)
for i in range(N):
x = np.random.rand(n)
y = 2.0*x - 1.0 + np.random.standard_t(df=2, size=n)
solL2 = optimize.minimize(fun=costL2, x0=[0, 0],
method='SLSQP', args=(x, y))
errL2[i] = np.abs(solL2.x[0] - 2.0)
solL1 = optimize.minimize(fun=costL1, x0=[0, 0],
method='SLSQP', args=(x, y))
errL1[i] = np.abs(solL1.x[0] - 2.0)
solLLog = optimize.minimize(fun=costLLog, x0=[0, 0],
method='SLSQP', args=(x, y))
errLLog[i] = np.sqrt((solLLog.x[0] - 2)**2)
print(" L2 mean:", np.mean(errL2))
print(" L1 mean:", np.mean(errL1))
print(" LLog mean:", np.mean(errLLog))
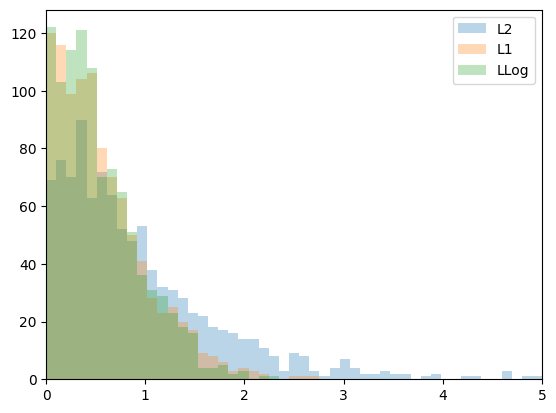
plt.figure(1)
plt.hist(errL2, bins=np.linspace(0,5,50),
label='L2', alpha=0.3)
plt.hist(errL1, bins=np.linspace(0,5,50),
label='L1', alpha=0.3)
plt.hist(errLLog, bins=np.linspace(0,5,50),
label='LLog', alpha=0.3)
plt.xlim([0, 5])
plt.legend()
plt.show()
L2 mean: 0.9933721689676688
L1 mean: 0.5684948908980173
LLog mean: 0.5416548165228569
# using white noise error
errL2 = np.zeros(N)
errL1 = np.zeros(N)
errLLog = np.zeros(N)
for i in range(N):
x = np.random.rand(n)
y = 2.0*x - 1.0 + np.random.rand(n) # white noise error
solL2 = optimize.minimize(fun=costL2, x0=[0, 0],
method='SLSQP', args=(x, y))
errL2[i] = np.abs(solL2.x[0] - 2.0)
solL1 = optimize.minimize(fun=costL1, x0=[0, 0],
method='SLSQP', args=(x, y))
errL1[i] = np.abs(solL1.x[0] - 2.0)
solLLog = optimize.minimize(fun=costLLog, x0=[0, 0],
method='SLSQP', args=(x, y))
errLLog[i] = np.sqrt(np.sum((solLLog.x[0] - 2)**2))
print(" L2 mean:", np.mean(errL2))
print(" L1 mean:", np.mean(errL1))
print(" LLog mean:", np.mean(errLLog))
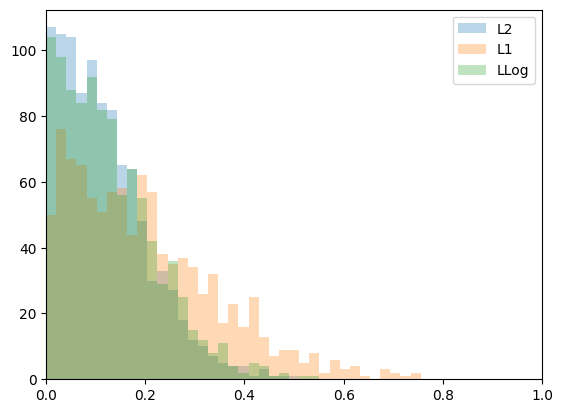
plt.figure(1)
plt.hist(errL2, bins=np.linspace(0,1,50),
label='L2', alpha=0.3)
plt.hist(errL1, bins=np.linspace(0,1,50),
label='L1', alpha=0.3)
plt.hist(errLLog, bins=np.linspace(0,1,50),
label='LLog', alpha=0.3)
plt.xlim([0, 1])
plt.legend()
plt.show()
L2 mean: 0.11838018024314924
L1 mean: 0.19834605402651811
LLog mean: 0.12783320665351436